Research Article
Antibacterial Activity of Cu Nanoparticles against E. coli, Staphylococcus aureus and Pseudomonas aeruginosa
Lalit Yadav 1, Ravi Mani Tripathi 1, Ram Prasad 2, Ramesh Namdeo Pudake 1, Jagjiwan Mittal 1*
1 Amity Institute of Nanoatechnology, Amity University, Noida-201301, Uttar Pradesh, India.
2 Amity Institute of Microbial Technology, Amity University, Noida-201301, Uttar Pradesh, India.
* Corresponding author. E-mail: jmittal@amity.edu
Received: Feb. 10, 2017; Accepted: Mar. 13, 2017; Published: Mar. 17, 2017.
Citation: Lalit Yadav, Ravi Mani Tripathi, Ram Prasad, Ramesh Namdeo Pudake, and Jagjiwan Mittal, Antibacterial Activity of Cu Nanoparticles against E. coli, Staphylococcus aureus and Pseudomonas aeruginosa. Nano Biomed. Eng., 2017, 9(1): 9-14.
DOI: 10.5101/nbe.v9i1.p9-14.
Abstract
Copper nanoparticles (CuNPs) were synthesized under ambient temperature by wet chemical method and evaluated their antibacterial activity by the means of well diffusion method. In this study, we have analyzed the effect of NaBH4 concentration on the synthesis of CuNPs. Surface plasmon resonance (SPR) was occurred at 575 nm. Scanning electron microscopy micrographs shows CuNPs have spherical morphology with size range of 20-40 nm. Dynamic light scattering (DLS) showed the particles size increases with the concentration of reducing agent NaBH4. As prepared, CuNPs, demonstrated antibacterial activity against E. coli, Staphylococcus aureus and Pseudomonas aeruginosa. The potential mechanism of antibacterial activity was discussed with diagrammatic representation.
Keywords: Copper; Nanoparticles; Particle size distribution; Electron microscopy; Spectroscopy; E. coli; Staphylococcus aureus; Pseudomonas aeruginosa; Antibacterial activity
Introduction
In chemical sciences, synthesis of transition metal and metal oxide nanoparticles (NPs) is a growing research field [1,2]. Copper nanoparticles (CuNPs) have attracted great interest due to their optical, catalytic, mechanical and electrical properties. Copper is a good alternative material for noble metals such as gold and silver, as it is highly conductive and much more economical than the preceding ones [3,4,5]. There are different methods of synthesis of CuNPs, including thermal reduction [6], metal-vapor synthesis [7], chemical reduction [8], vacuum vapor deposition [9], radiation methods [10], micro emulsion techniques [11], polyol processes [12] and laser ablation [13]. It has been reported that metal nanoparticles (Ag, Cu, CuO, Au) exhibit a wide spectrum of antimicrobial activity against different species of microorganisms [14,15]. The antibacterial activity of silver nanoparticle was analyzed against gram positive and gram negative bacteria, which revealed silver nanostructured material is a prominent source for antibacterial activity [16,17]. Antibacterial activity of CuNPs has been reported against Escherichia coli [18,19] and Staphylococcus aureus [20,21]. However, there are some concerns related to environmental and human safety regarding the release and consumption of metal nanoparticles, which are yet to be explored. Excessive release of silver, for example, causes environmental pollution which in turn makes silver harmful to humans and animals. Copper is no exception, because an excess of copper in the human body leads to the generation of the most damaging radicals, such as the hydroxyl radical [22]. However, there are copper-transporting adenosine triphosphatases (Cu-ATPases), including ATP7A and ATP7B, which play an important role in the homeostasis and export,of excess copper through the intestine (ATP7A) as feces, the liver (ATP7B) as a bile product, and the mammary gland (ATP7B) as milk [22,23]. In the present study, we synthesized copper nanoparticles under ambient conditions using simple and rapid wet chemical method. These nanoparticles were studied for their antibacterial activities against strain of E. coli, Staphylococcus aureus and Pseudomonas aeruginosa. The bacterial strain selection was done by means of different gram strain and in terms of their clinical significance. A possible mechanism of antibacterial activities of CuNPs is proposed with schematic presentations in the paper.
Materials and Methods
Chemicals and bacterial strains
Copper sulfate (CuSO4.5H2O), polyvinyl alcohol (PVA), sodium borohydride (NaBH4) and sodium chloride (NaCl) were used as received. Commercially available Mullar Hinton media was used in the study for in-vitro growth of bacteria. The strain of E. coli, Staphylococcus aureus and Pseudomonas aeruginosa were obtained from Amity Institute of Microbial Technology, Amity University Uttar Pradesh, India.
Synthesis and characterization of CuNPs
Copper nanoparticles were synthesized by method used in previous study (Deng et.al., 2011) with some modifications. Briefly, 0.034 M copper sulfate (CuSO4. 5H2O) solution was mixed with 2.5% polyvinyl alcohol solution in Erlenmeyer flask. Subsequently, 0.5 Molar solution of sodium borohydride (NaBH4) was added in above solution drop by drop with vigorous stirring. The formation of CuNPs was monitored using Double Beam UV visible spectrophotometer in 1 cm path length quartz cuvette by scanning the sample between 400 to 700 nm. The size distribution profile of synthesized CuNPs was analyzed by dynamic light scattering (DLS). The size and morphology of synthesized nanoparticles were analyzed using scanning electron microscopy (SEM) and high resolution transmission electron microscopy (HRTEM). Samples for SEM and TEM were prepared by drop and dry method. Few drops of liquids containing Cu nanoparticles were put on the SEM sample holder and TEM grid, and dried at 50 °C.
Antibacterial analysis of CuNPs
The antibacterial activity of chemically synthesized CuNPs was evaluated against E. coli, Staphylococcus aureus and Pseudomonas aeruginosa by well diffusion method [14,16]. Luria-Bertani (LB) media were used for the growth of bacterial strains. After solidification of LB media, 5 mm diameter wells were punched with the help of cork borer. Uniform Spreading of bacterial culture was done on each agar plates. In every well, different concentrations of CuNPs, viz 20, 40, 60 and 80μL were added and plates were incubated at 37 oC for 24 hours. After incubation, the zone of inhibition was measured. Each experiment was repeated in triplicates.
Results and Discussion
UV-Vis Spectroscopy analysis
The absorption spectrum between 400 to 700 nm in UV-Vis spectroscopy for the prepared nanoparticles is shown in Fig. 1. A broad band is observed between 550 and 600 nm with a peak at 575 nm in the yellow region of the visible range of light spectrum. General surface plasmon resonance of CuNPs was observed in the range of 569- 597 nm in earlier studies [24]. Therefore, the presence of a band in this wave length range in the present study confirms the formation of CuNPs.
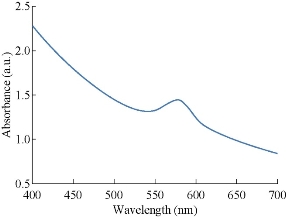
Fig. 1 UV-Vis study of the prepared CuNPs shows a peak at 580 nm.
Dynamic light scattering (DLS) analysis
Dynamic Light Scattering (DLS) was used to determine the size distribution profile, the poly-dispersities and the aggregation effects of colloidal samples. The copper nanoparticles were synthesized in different sizes using various concentrations of NaBH4 solution in the reaction mixture. As shown in Fig. 2, the color of colloidal solutions darkens with the increasing concentration of reducing agent. Fig.3 (a)-(d) shows the change in sizes with the amount of NaBH4. As shown in Fig. 3, the diameter of nanoparticles increased with the increasing concentration of NaBH4 solution. The diameter increased from 13.5 nm to 190.14 nm. The effect of reducing agent was easily identified on the basis of color of colloidal solution of copper nanoparticles. Diameter increase was also accompanied with the change in the color of particles. As revealed by DLS study (Fig. 3(a)), nanoparticles in the transparent solution as shown in Fig. 2(a) have a comparatively smaller average size of 13.54 nm than the nanoparticle synthesized by using more concentrations of NaBH4 58.78, 58.78, 105.7 and 190.14 nm, respectively (Fig 2(b)-(e)).
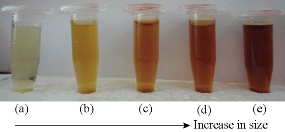

Fig. 2 Size measurement of CuNPs using differential light scattering after adding different amounts of NaBH4.
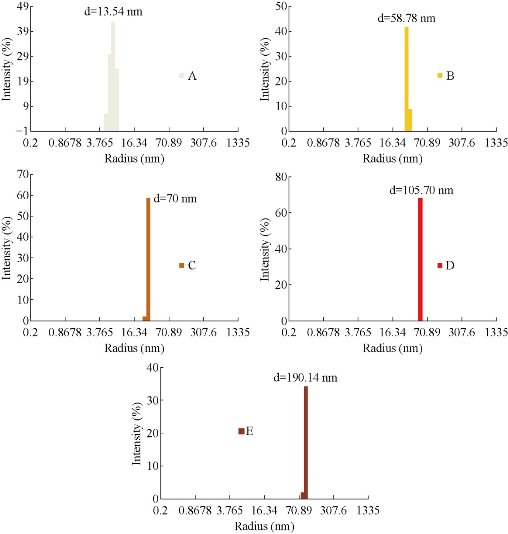
Fig. 3 Change in color of the CuNPs with increase in diameter after treatment with NaBH4.
Scanning electron microscopy (SEM) analysis
SEM micrographs in Fig. 4 under magnification of 92.35K show the Cu nanoparticles of 100-150 nm. However, high magnification study of the sample using 203.53K revealed existence of much smaller nanoparticles in a cluster.
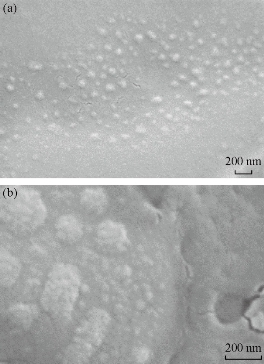

Fig. 4 SEM micrographs of CuNPs using magnifications of (a) ×92.35K; and (b) ×203.53K.
HRTEM study
HRTEM micrograph in Fig. 5(a) shows the two clusters of CuNPs nearby each other. The observation of a portion of a cluster in high magnification in Fig.5(b) shows the very small sized nanoparticles of 4-5 nm diameters. HRTEM study confirmed that the nanoparticles seen in the SEM is the cluster of CuNPs. EDX study of the portion demonstrates the presence of Cu with slight amount of oxygen in these nanoparticles.
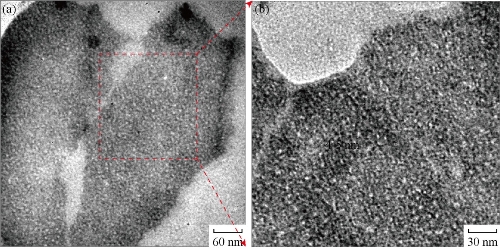
Fig. 5 HRTEM micrographs of (a) two clusters of CuNPs; and (b) a portion of a cluster in (a) in high magnification.
Antibacterial activity analysis
Chemically synthesized copper nanoparticles were analyzed for antibacterial activity against bacteria such as E. coli, Staphylococcus aureus and Pseudomonas aeruginosa. These tests were performed on LB plates by well diffusion method, and the growth of bacteria was inhibited by CuNPs which formed an inhibitory zone as shown in Fig. 6. The highest zone of inhibition was observed for E. coli (Fig. 6 & 7) and had more average diameter of zone of inhibition as compared to other bacteria used (Fig. 6 & 7). Recently, there have been few reports on the mechanism of bactericidal activity of CuNPs when used alone or along with some other materials. The CuNPs has been found to generate reactive oxygen species (ROS) and to ultimately result in cell death. The size of the nano particle has also been found to be responsible for the toxicity; the smaller the size is, the greater toxicity it has. Bacteria are protected by a layer called cell wall - an essential element for its survival. It is composed of polysaccharides and peptides which are called peptidoglycan. The thickness of the cell wall varies in both gram-negative (thin layer of peptidoglycan) and gram-positive bacteria (thick layer of peptidoglycan). The copper nanoparticles directly bind with cell wall and interact with the membrane proteins, causing the perforation of membrane which results in the release of intracellular material [25]. In one report, CuNPs was found to act as an efflux inhibitor in bacterial cells, the antibacterial effect of which was partly mediated by particle effect and partly by ion effect [26]. Some other studies suggested that Cu ions originating from the NPs may interact with phosphorus and sulfur-containing biomolecules, such as DNA and protein to distort their structures and thus disrupt biochemical processes [27-30]. Thus, from the results of earlier and our studies, we propose that CuNPs prepared here cause bacterial cell death due to multimodal action (Fig 8).
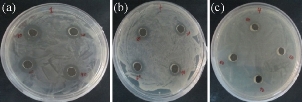
Fig 6 Antibacterial analysis against pathogenic bacteria: (a) E. Coli; (b) Staphylococcus aureus; and (c) Pseudomonas aeruginosa.
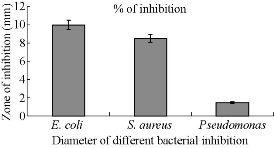
Fig. 7 Antibacterial activity of CuNPs against pathogenic bacteria in terms of zone of inhibition.
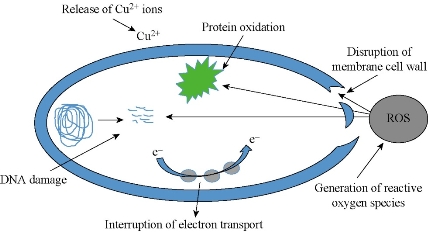
Fig. 8 Schematic representation of probable antimicrobial mechanism of CuNPs.
Conclusions
In this study, we synthesized copper nanoparticles with various sizes at ambient condition and characterized them with UV-Vis spectroscopy, DLS and SEM. The concentration of NaBH4 plays an important role in the size of nanoparticles. DLS shows that nanoparicles suspension color was darkening with the increasing concentration of NaBH4. SEM micrograph reveals that particles are spherical with 20- 40 nm in size. The synthesized nanoparticles shows effective antibacterial activity against E. coli, Staphylococcus aureus and Pseudomonas aeruginosa. The antibacterial mechanism was discussed with diagrammatical representation.
Acknowledgements
All authors are grateful to Amity University, Noida for providing excellent facility for above research work.
Conflict of Interest
The authors declare that they have no conflict of interest in the publication
References
Copyright© 2017 Lalit Yadav, Ravi Mani Tripathi, Ram Prasad, Ramesh Namdeo Pudake, and Jagjiwan Mittal. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.